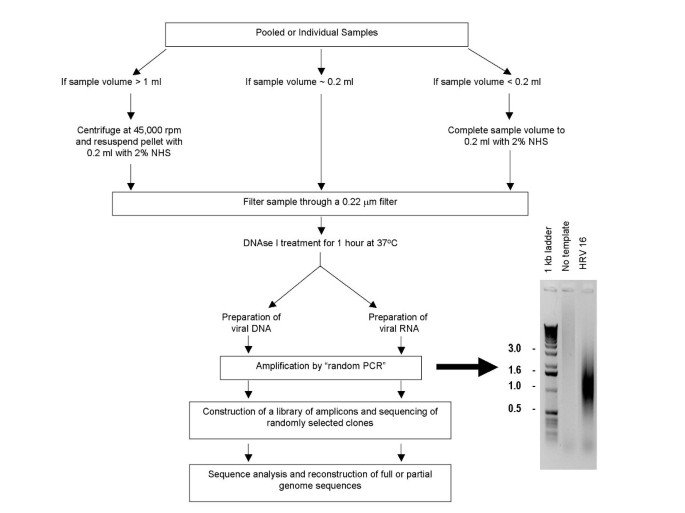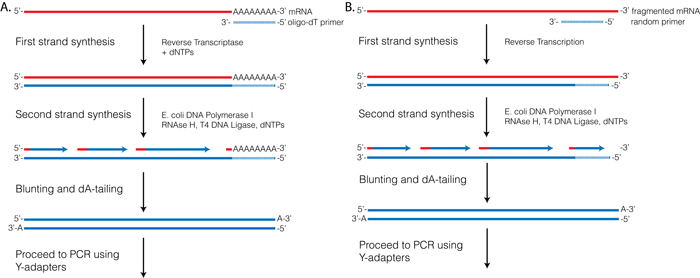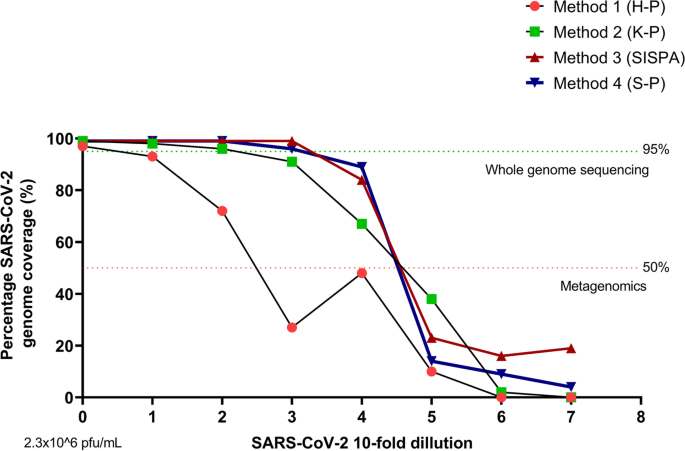
A random priming amplification method for whole genome sequencing of SARS-CoV-2 virus | BMC Genomics | Full Text
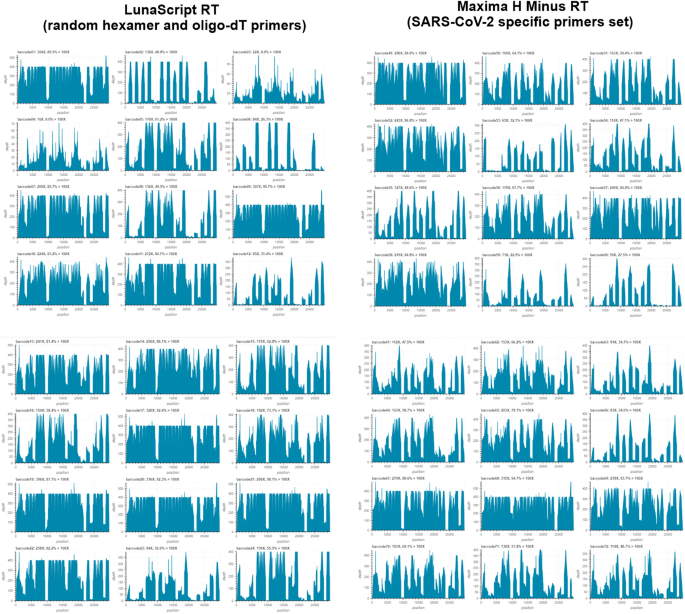
Universal whole-genome Oxford nanopore sequencing of SARS-CoV-2 using tiled amplicons | Scientific Reports

Protocol for High-Resolution Mapping of Splicing Products and Isoforms by RT-PCR Using Fluorescently Labeled Primers - ScienceDirect
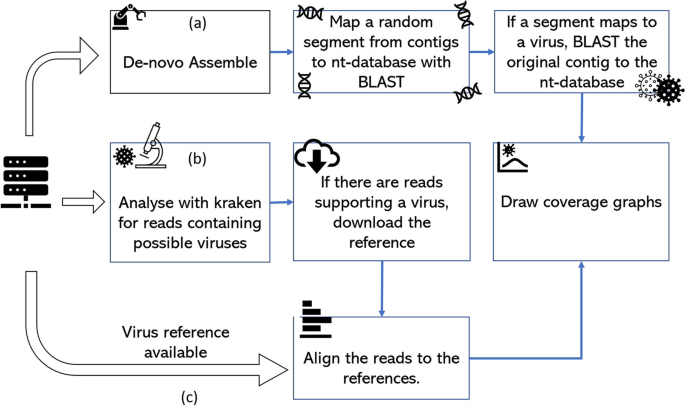
A random priming amplification method for whole genome sequencing of SARS-CoV-2 virus | BMC Genomics | Full Text
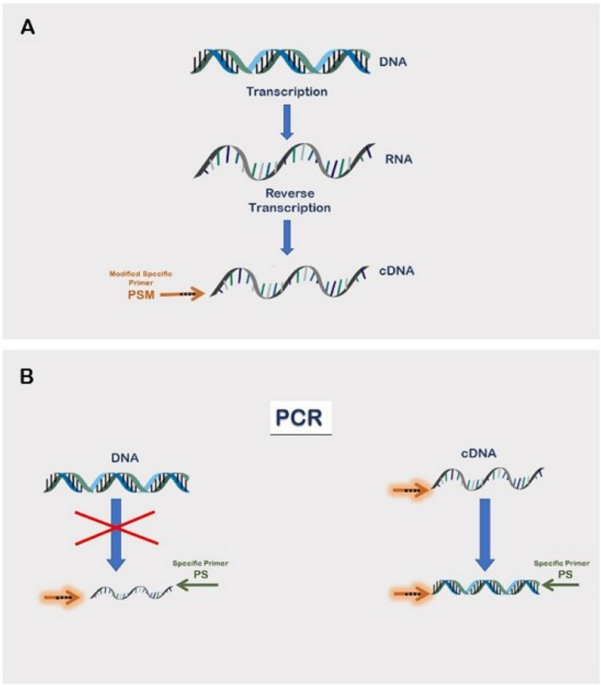
Reverse transcription-quantitative PCR (RT-qPCR) without the need for prior removal of DNA | Scientific Reports
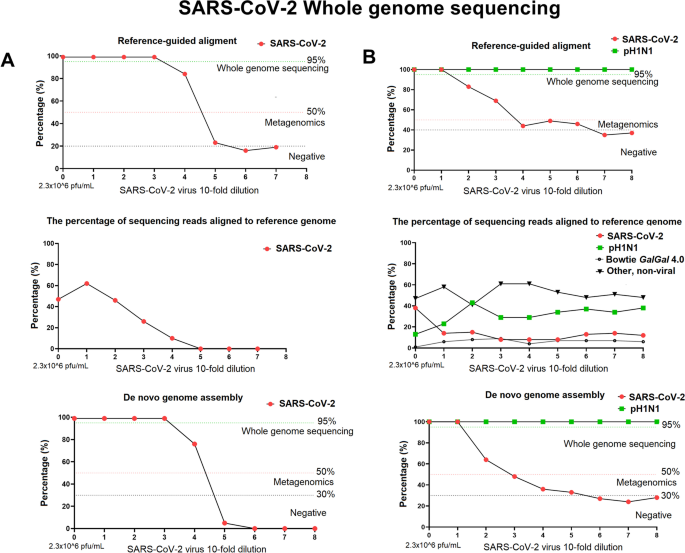
A random priming amplification method for whole genome sequencing of SARS-CoV-2 virus | BMC Genomics | Full Text

Use of Sequence-Independent, Single-Primer-Amplification (SISPA) for rapid detection, identification, and characterization of avian RNA viruses - ScienceDirect
Frontiers | In situ RT-PCR Optimized for Electron Microscopy Allows Description of Subcellular Morphology of Target mRNA-Expressing Cells in the Brain
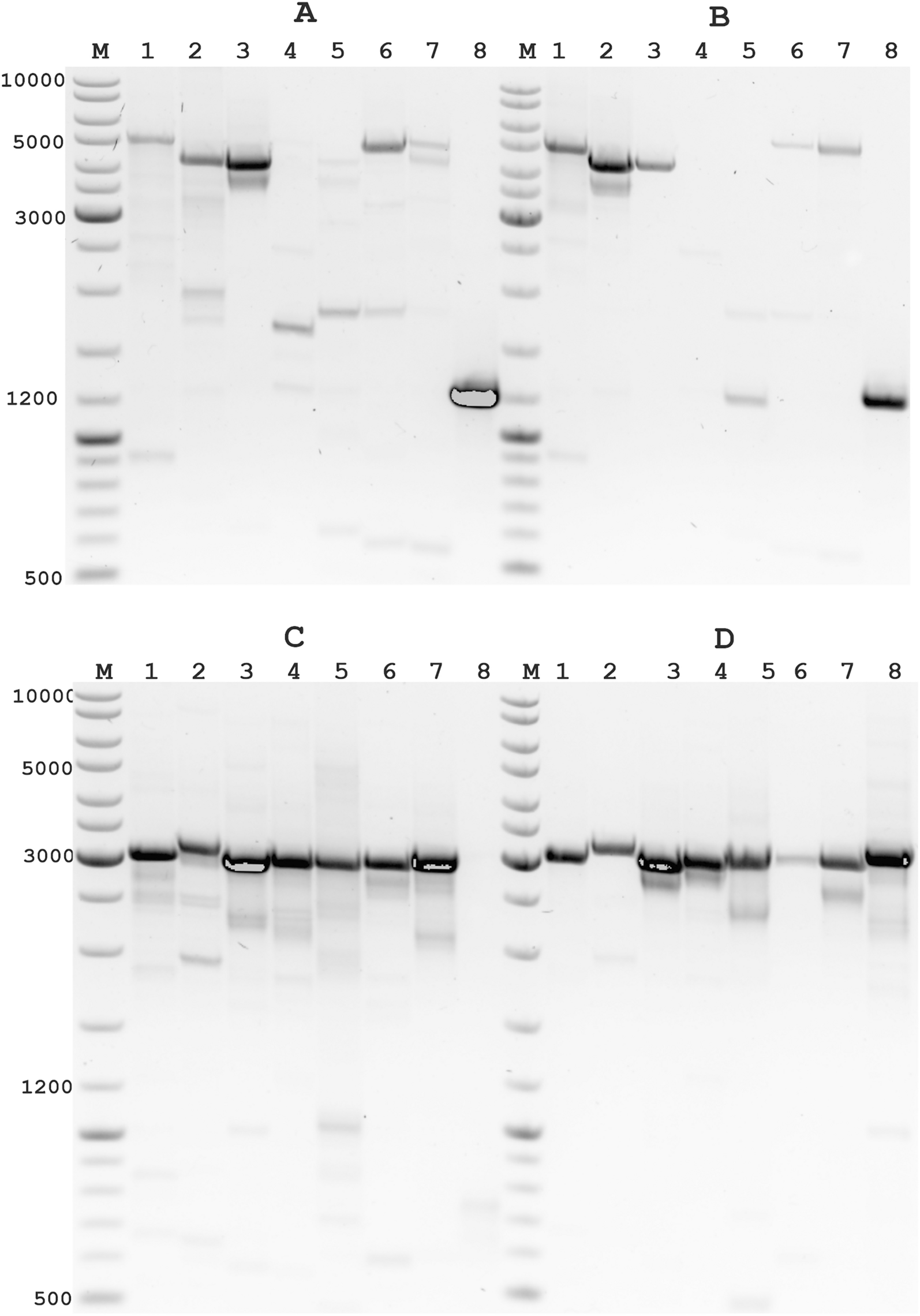
Universal whole-genome Oxford nanopore sequencing of SARS-CoV-2 using tiled amplicons | Scientific Reports
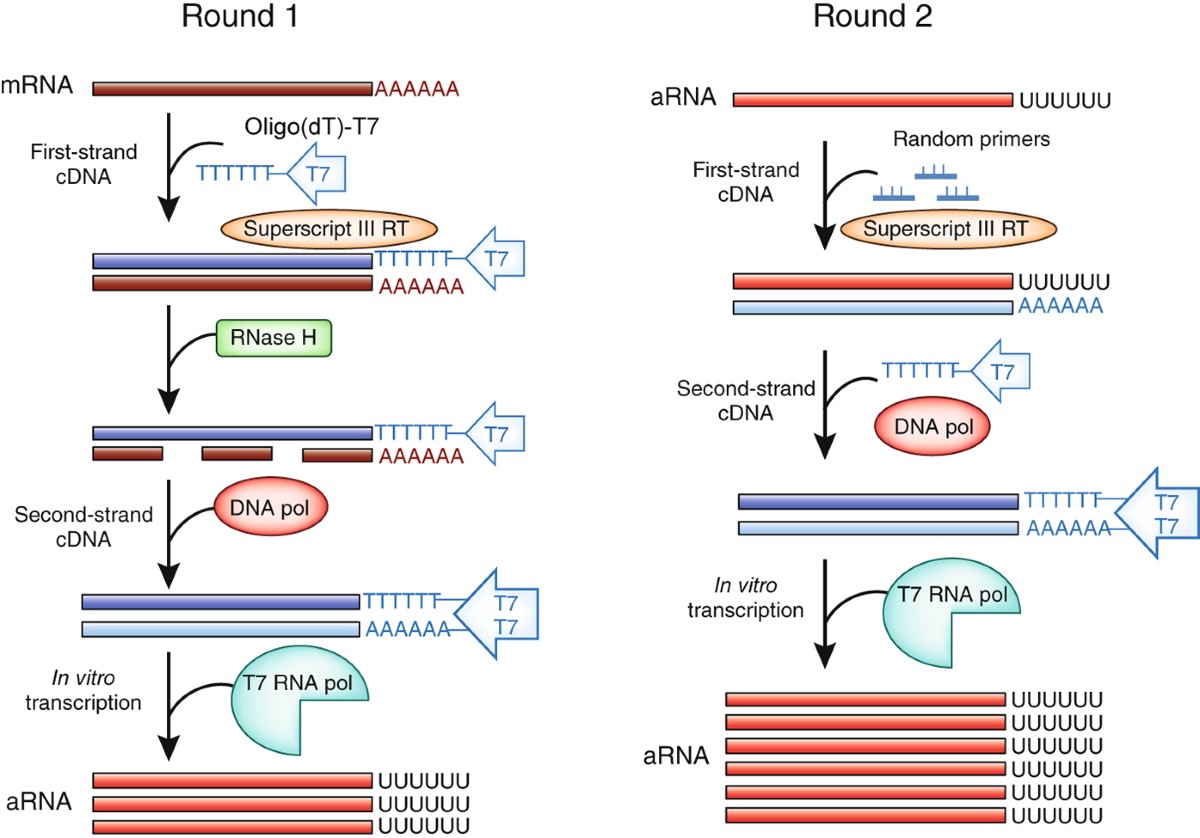
The successes and future prospects of the linear antisense RNA amplification methodology | Nature Protocols